Hyperscanning series part 5:
How to analyze a hyperscanning dataset with BrainVision Analyzer 2
by Dr. Bastian Schledde
Scientific Consultant (Brain Products)
![]() Have you been following our article series on hyperscanning? Maybe you have already recorded data from multiple participants and would like to go forward? Then you might be interested in the following tips around the analysis of a hyperscanning dataset.
Have you been following our article series on hyperscanning? Maybe you have already recorded data from multiple participants and would like to go forward? Then you might be interested in the following tips around the analysis of a hyperscanning dataset.
Getting started
 Hyperscanning research has been developed to study the brain in a social environment. The interactive and social nature of the human brain is largely omitted in conventional paradigms. This is a crucial limitation for research subjects such as social behavior, interpersonal coordination, interactive decision-making, or affective communication.
Hyperscanning research has been developed to study the brain in a social environment. The interactive and social nature of the human brain is largely omitted in conventional paradigms. This is a crucial limitation for research subjects such as social behavior, interpersonal coordination, interactive decision-making, or affective communication.
Hyperscanning allows brain activity from two or more people to be observed simultaneously. Researchers are aiming to find inter-brain correlations that reflect upon interpersonal dynamics. For example, if two participants coordinate a simple, self-initiated button press, coherence between frontal regions of both brains increases [1].
The methods for inter-brain correlations are mostly the same as used for intra-brain analysis. Despite its young age, the field already provides recommendations for hyperscanning analysis and tools [2,3]. BrainVision Analyzer 2 provides some of these methods in the Connectivity modules and, leaving aside methodological discussions, here we provide some recommendations for implementing a hyperscanning pipeline in Analyzer.
Multi-participant vs. single-participant processing
The recording setups for hyperscanning studies can be manifold. This is also reflected in the variety of hardware solutions that we have presented in our recent articles.
As a result, data recorded from multiple participants can live in multiple or single files. For example, dyadic recordings typically result in a single recording file while setups with more participants or mobile systems may result in multiple files.
Generally, transformations in Analyzer process only channels from one History Node, with few exceptions. Connectivity modules need channels of all participants to be in the same dataset to calculate correlations among them. When recording multiple files or splitting datasets into participants, this is where all channels need to be merged into one dataset.
Figure 1 outlines different pathways starting from Raw Data files loaded into the Analyzer workspace, encompassing preprocessing until connectivity analysis.
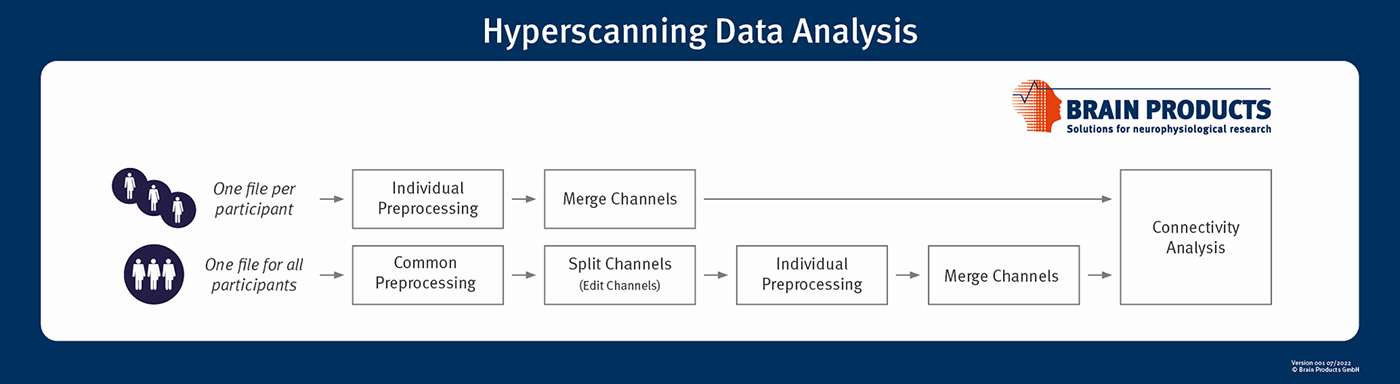
Figure 1: Example processing pathways for single and multiple Raw Data files. Edit Channels transformation allows datasets to be split into individual participants. Merge Channels Solution can merge data back together for correlations between participants. We recommend to merge datasets only if they are sufficiently synchronized.
If channels of all participants are recorded into one file, it makes sense to split the dataset for processing steps applicable only to individual participants. Our suggestion is to split the set of channels once using the Edit Channels transformation and disabling channels of all other participants. Of course, Analyzer also offers a channel selection option in most transformations, but this requires repeated manual selection for many steps in the pipeline.
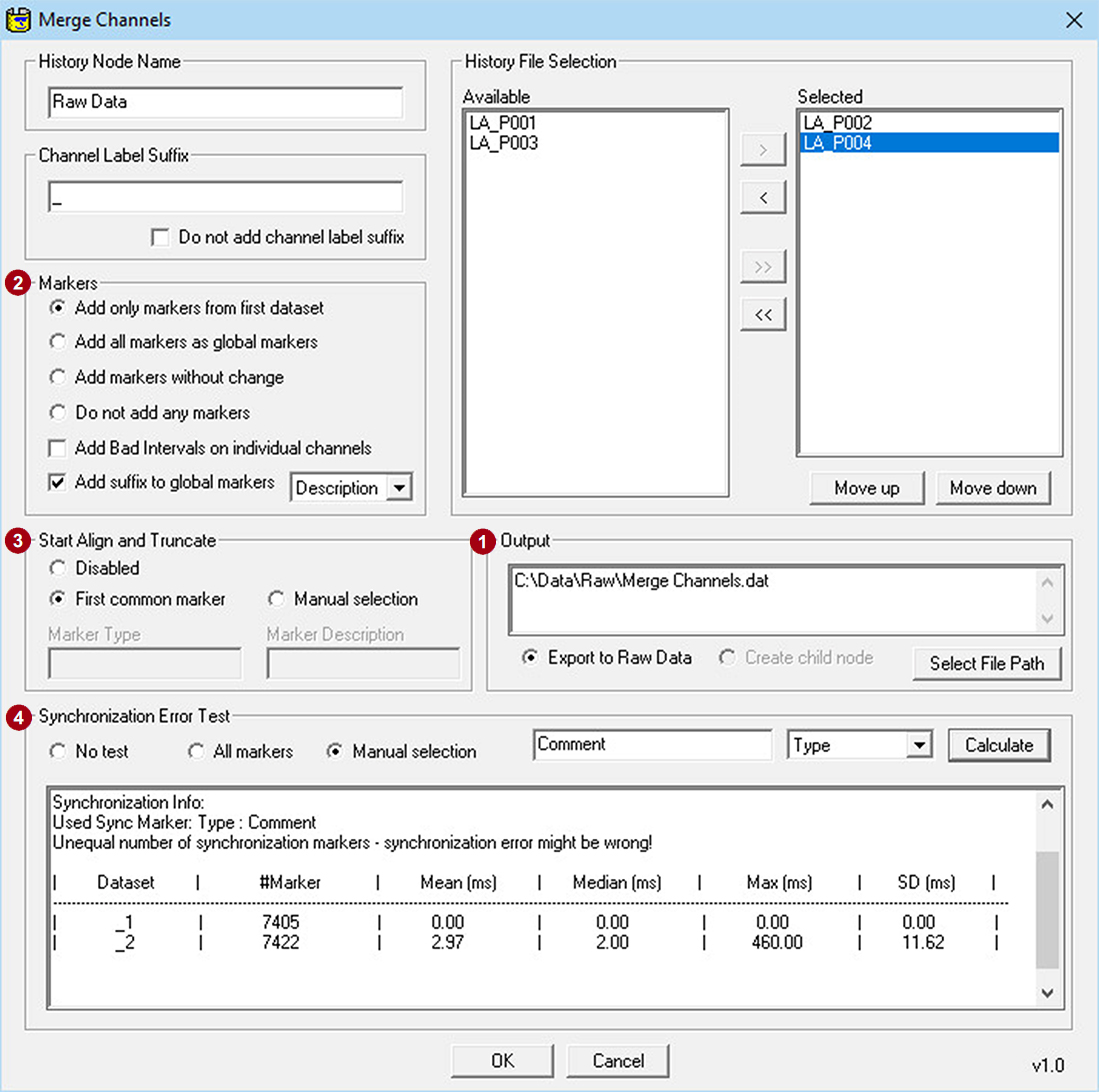
The Merge Channels Solution allows merging sets of channels together before correlational analysis. Note that datasets that shall be merged are recommended to be synchronized. If the acquisition setup is stable, markers and data are recorded without loss or jitter, correcting clock-offsets and drifts might be sufficient. Let us know if you want us to help with a workaround in Analyzer to achieve it.
Together, the Edit Channels transformation and the Merge Channels Solution offer flexible, complimentary data handling options that meet many user needs.
What to consider during preprocessing
Individual Preprocessing Steps
It is important to consider which steps are part of preprocessing and whether it is okay to apply them on the combined set of channels or only on single participant data. For example, if you plan to use the common average reference you might want to use only channels from the same participant for it. The same might be true for computing independent components. Other processing steps are less restrictive such as changing the sampling rate or filtering data.
Artifact Handling
In this context, it is worth looking at artifact handling too. The question is whether an artifact in a single participant must lead to the rejection of data for all participants in the dataset. It is necessary when correlating this participant with another one. It may not be necessary, when another participant of the same dataset is analyzed independently, then it leads to unnecessary loss of data.
In Analyzer, there is a neat solution to optimize rejection and inclusion of data. Noisy intervals can be marked with artifact-handling transformations during preprocessing on the individual participant level. The data is not rejected at this point but only marked as a “Bad Interval”.
When merging individual participant datasets, Bad Interval makers can be assigned to channels of all participants or only to participants where the artifact occurred. In some transformations, such as the Average or Cross-Correlation, segments can be excluded only for channels of the affected participant. In other transformations, such as the Coherence or Correlation Measures, segments from all channels will be excluded irrespective of whether Bad Intervals are assigned to single participants or all channels of the dataset.
Channel Labels and Coordinates
When recording participants in one file or after merging, a dataset can contain duplicate channel labels and coordinates. Analyzer tries to avoid this situation and does not allow it in many transformations. For example, you cannot use the IIR Filters transformation if duplicated channel labels exist. Duplicated channel coordinates can lead to warnings in views, or even prevent the usage of transformations that rely on channel coordinates such as the Current Source Density. While some transformations may be applied to such a dataset, it is generally easier to work with channels only from one head. Therefore, we recommend splitting the dataset into individual participants using the Edit Channels transformation early in the processing pipeline.
The Merge Channels Solution offers a dedicated feature to add channel label suffixes, for example “Fp1_1” when merging them back together to prevent duplicated channel labels.
If you need to assign default channel coordinates, for example for 10-20 labels, you will also profit from having individual participant datasets. 10-20 electrode positions can be assigned automatically with the Edit Channels transformation. However, this works only if standard electrode labels that do not have a participant suffix are used. We created an electrode coordinate file for you in case you need to assign 10-20 coordinates to channels with a suffix. It works for two participants if an underscore suffix is used, for example “Fp1_1” and “Fp1_2”. Feel free to download this file and assign electrode coordinates via the option “Load Positions from File” in the Edit Channels transformation or by adding the file to your default electrode positions via File > Preferences > Electrodes.
All articles of the hyperscanning series
Merge Channels Solution
Merge Channels Solution merges datasets from different History Files or from different History Nodes of the same History File. If you have recorded individual Raw Data files for each participant, you would be merging from different History Files (Figure 2A). Otherwise, if you have all participants in the same Raw Data file, but different History Nodes, you would be merging within a History File (Figure 2B). In this case you have the option to create a new child node below the node of the active dataset. This node can be used in a History Template, making it easy to apply a processing pipeline, for example to many dyadic recordings.
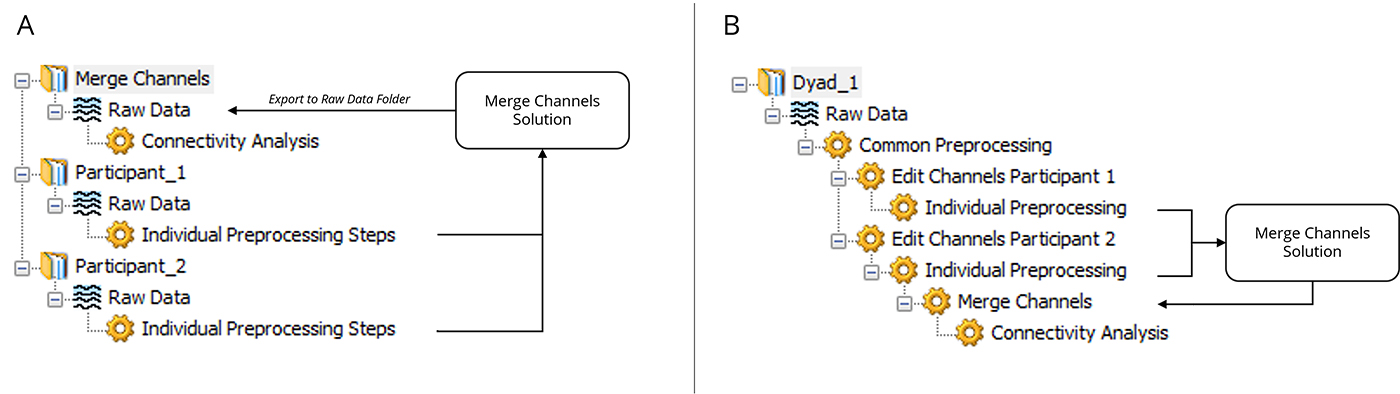
Figure 2: Two modes of merging. A: When data is recorded in multiple files, datasets can be merged at any point during processing, before or after individual preprocessing steps. B: When data of all participants is recorded into one file, the datasets might be split into participants during preprocessing. The Merge Channels Solution can merge datasets into one before connectivity analysis.
More features:
A few words about the marker handling options ():
How to do inter-brain connectivity
Hyperscanning research can be informed by intra- and inter-brain analysis. Intra-brain analysis looks at the individual participant and quantifies, for example, the ERP, spectral power, or connectivity. Inter-brain analysis is interesting because it measures whether brains of different participants engage in similar processes during social interaction. This is often quantified with connectivity methods that are also applied for intra-brain analysis.
Similarity between brain activity of two participants is measured by correlating the signal amplitudes, phases, or both. Which approach is more suitable depends on the requirements of the research question. Amplitude based measures include the correlation of power or envelope. You can obtain them in Analyzer with the FFT, Complex Demodulation and Correlation Measures transformations. The Complex Demodulation can be understood as the analytical signal comparable to the Hilbert transform.
More common, though, are Fourier spectrum or Wavelet-based metrics. For example, the Phase Locking Value (PLV) estimates phase-synchrony while the Magnitude-Squared Coherence calculates similarity based on phase and amplitude. These measures are available in Analyzer via the FFT, Wavelet, Coherence or Correlation Measures transformations. Check out our webinar recording on Phase- and Connectivity Analysis and our newsletter article including a description for implementing the PLV. Additional methods for quantifying correlation and other comparisons are offered in the Cross-Correlation and Data Comparison transformations.
Analyzer has dedicated Channel Pair views for displaying relations between channels. They are designed for single heads and show connectivity between channels of the same participant. If you are comparing across channels and participants, this display might make sense to some degree. It does not make sense, though, if you are comparing the same channels between participants. Instead, we recommend displaying the frequency spectrum or time (-frequency) domain signals in the Grid View.
Pairwise correlations as in the above-mentioned connectivity modules can get quite complex. This happens, in particular, if pairs are formed between multiple channels of more than two subjects. Entering or editing many channel pairs in the Manual Channel Pair Selection is quite cumbersome. For correlating symmetric pairs between two participants with conventional 10-20 labels and channel suffixes (as described above), we have created a BrainVision Graph file for your convenience. You can load it in the Manual Channel Pair Selection wizard via Load Graph in transformations dealing with channel pairs.
We recommend, however, to consider reducing complexity for computational reasons as well. You can, for example, choose a subset of representative channels to perform pairwise correlations.
Once you are happy with your analysis, you can export data for statistical evaluation. If you have been looking at time-domain or frequency domain data, you can use the Area Information or Peak Information Export modules. If you have calculated time-frequency domain metrics, we recommend using the Wavelet Data Export Solution allowing you to aggregate and export data from a time and frequency range.
Concluding remarks
Irrespective of how you have recorded your data, we hope the tools above help to pursue your hyperscanning analysis as you desire.
History Files in Analyzer can get quite large if the dataset is large. Sometimes this leads to memory issues. If your paradigm contains many participants or is rather long, we recommend considering having participants in individual but synchronized Raw Data files. Reducing the size of data before merging, for example by segmentation, can help managing memory bottlenecks.
While having all participants in one dataset is necessary for correlation, for other processing steps you might benefit from handling one dataset at a time. For example, when screening for artifacts. In other situations, the method might even be invalidated if all participants are in one dataset. For example, if you want to use the Mapping View or compute source activity with the LORETA transformation. It makes sense to plan out your analysis thoroughly before starting to record such a complex dataset.
Make sure to read along your peer literature to understand the pitfalls accompanying hyperscanning connectivity findings. One caveat to consider is whether the modulation you observe relates to the interactive aspects of your paradigm or to a common external stimulus. The good news is that other caveats such as volume conduction or the common reference problem are not issues for inter-brain connectivity.
As usual we are more than happy to help you in every step of your analysis. Please do not hesitate to write us an email via support@brainproducts.com.
References
[1] Czeszumski, A., Eustergerling, S., Lang, A., Menrath, D., Gerstenberger, M., Schuberth, S., Schreiber, F., Rendon, Z. Z., & König, P. (2020).
Hyperscanning: A Valid Method to Study Neural Inter-brain Underpinnings of Social Interaction.
Frontiers in human neuroscience, 14, 39. https://doi.org/10.3389/fnhum.2020.00039.[2] Ayrolles, A., Brun, F., Chen, P., Djalovski, A., Beauxis, Y., Delorme, R., Bourgeron, T., Dikker, S., & Dumas, G. (2021).
HyPyP: a Hyperscanning Python Pipeline for inter-brain connectivity analysis.
Social cognitive and affective neuroscience, 16(1-2), 72–83. https://doi.org/10.1093/scan/nsaa141[3] Kayhan, E., Matthes, D., Marriott Haresign, I., Bánki, A., Michel, C., Langeloh, M., Wass, S., & Hoehl, S. (2022).
DEEP: A dual EEG pipeline for developmental hyperscanning studies.
Developmental cognitive neuroscience, 54, 101104. https://doi.org/10.1016/j.dcn.2022.101104