Non-parametric permutation statistics made easy – Analyzer 2 and BESA Statistics 2.0
by Michael Hoppstädter
Scientific Consultant (Brain Products)
![]() Would you like to statistically explore all time-points, all electrodes for all experimental conditions in less than one hour? BESA Statistics is an easy to use stand-alone application to calculate non-parametric permutation statistics for EEG, MEG and source imaging data. The newest version 2.0 released in July 2015 is directly compatible with the BrainVision native format which allows you to do all your data processing in BrainVision Analyzer 2 and then switch to BESA Statistics for statistical evaluation of your neurophysiological data.
Would you like to statistically explore all time-points, all electrodes for all experimental conditions in less than one hour? BESA Statistics is an easy to use stand-alone application to calculate non-parametric permutation statistics for EEG, MEG and source imaging data. The newest version 2.0 released in July 2015 is directly compatible with the BrainVision native format which allows you to do all your data processing in BrainVision Analyzer 2 and then switch to BESA Statistics for statistical evaluation of your neurophysiological data.
The following article should give you an overview on the statistical procedures used in BESA statistics, how the data is handled and finally a recipe what you have to do in Analyzer 2 to export your data to BESA Statistics.
Statistical features of BESA Statistics 2.0
BESA Statistics works mainly in a two-step fashion. The statistical procedures comprise a preliminary parametric t-test, F-test, or correlation, to derive significant data clusters that are subsequently tested with permutation statistics. The initial parametric statistics are calculated separately for each time (or time-frequency) sample and for all channels. This massive univariate testing inflates the degrees of freedom for the statistical test and thus leads to the well-known problem of alpha error accumulation due to multiple comparisons. The permutation test in contrast is parameter-free and not calculated sample-wise. In each permutation step, subjects or conditions are re-shuffled, statistically significant adjoining samples are clustered, and cluster values are used to build a parameter-free distribution. From the resulting distribution of permuted cluster values, it is then possible to derive the exact probability of the clusters found in the original datasets. This statistic is no longer subject to the multiple comparisons problem (see Maris and Oostenveld, 2007 for an in-depth overview on permutation statistics as implemented in BESA Statistics). For designs with more than two groups / conditions, it is also possible to calculate post-hoc pairwise comparisons between group / condition levels. For this purpose, a permutation statistic based on Scheffé’s test is calculated for each pairwise comparison. To counteract the alpha error accumulation due to multiple pairwise tests, a Bonferroni-Holm correction is applied.
Data handling: Projects and Workflows
The first step in the statistical analysis sets constraints on the statistical test design (test between groups or conditions, number of groups / conditions, etc.). The user decides on a project type which can be either t-test, ANOVA (analysis of variance) or correlation.
The second decision is then on the data type. BESA Statistics allows for evaluation of different data types (ERP or ERF, time-frequency data, source waveforms or distributed source images) from different measurement modalities (EEG, MEG, intracranial data or polygraphic data). After project and data type were defined (the so-called Project Targets), the data files are loaded and ready for statistics. If you want to load data in BrainVision native format, you can set data type to BVA Files (*.vhdr) in the search mask.
The whole sequence of processing steps is organized in a Workflow which makes it easy to switch between different calculations or to jump back and change parameters (see Figure 1).
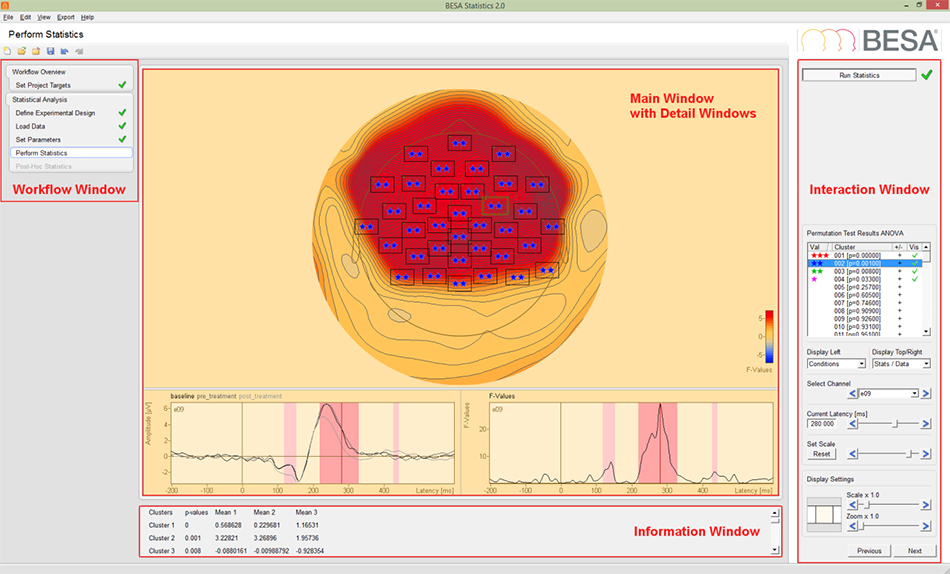
Figure 1: Main Interface of BESA Statistics 2.0. The workflow can be used to jump back and forth between individual steps. Channels and time (or time-frequency) ranges belonging to significant clusters are marked by asterisks and red overlays, respectively. Statistical values are listed in the Information Window and can be exported.
Interfacing between Analyzer 2 and BESA Statistics 2.0
The new version 2.0 of BESA Statistics offers full compatibility with BrainVision native format for time and time-frequency domain data. The best way to generate export files in this format is to use the Generic Data Export. In a few easy steps you can generate a new set of data, header and marker files. For a more extensive introduction into how to use the Generic Data Export function in general, please have a look at the article “Exports for all occasions – A selective overview of Analyzer 2’s most useful export options”.
For interfacing with BESA Statistics 2.0
… it is important that you consider the following points when you export your averaged data from Analyzer 2:
- Write header and marker files in text format (.vhdr, .vmrk)
- Check that the extension of the data file is set to .dat
- Ensure that the data file is binary and use PC format for line delimiters
- Data has to be exported in 32 bit floating point format
- Export all available EEG channels for statistics to achieve dense spatial clustering
If you want to calculate statistics on time-frequency domain data,
… there are some additional points to consider in your Wavelet transformation in Analyzer 2:
- Time-frequency statistics in BESA Statistics is only for use with continuous wavelets (Morlet, Morlet Complex, Mexican Hat)
- Wavelet layers can be linearly or logarithmically spaced, but it is generally recommended to use logarithmic scaling
- All output values are supported (complex or real-valued)
Importantly, if you use the Generic Data Export to get your data from Analyzer 2 into BESA Statistics 2.0, it is not necessary to provide an additional electrode positions file. The position data for spatial clustering and mapping visualizations will be read directly from the header file.
Concluding remarks
BESA Statistics 2.0 is a user-friendly statistics program that generates, without much manual interaction, nice graphics and statistical values optimized for publications – and delivers this without having to worry about multiple comparisons thanks to cluster permutation tests. Given the new compatibility with the BrainVision native format, BESA Statistics can be considered as a powerful addition to your data processing in Analyzer 2. For more information, please visit BESA’s webpage at http://www.besa.de/products/besa-statistics. All questions about the use of BESA Statistics can be directly addressed to BESA Support via their website.
Reference
Maris E, and Oostenveld R.
Nonparametric statistical testing of EEG- and MEG-data.
J Neurosci Meth 164: 177-190, 2007.

